This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Protein Homology
Homology is an indication of how similar two protein sequences compare to each other. Generally, proteins will be more similar to each other when they perform similar functions and have more recent common ancestors. Homology can be an important tool in the study of proteins because it can indicate which proteins are more important than others across species and what specific amino acid residues are necessary for the proteins to function correctly. It can also be used to study disease, when changes in the amino acid sequence of a protein cause a similar phenotypic response in many species [1].
The human SLC6A4 protein can be described as follows:
The human SLC6A4 protein can be described as follows:
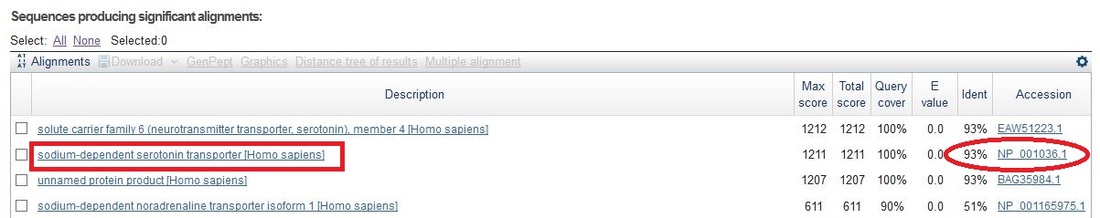
Homo sapiens (Human)
Solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4)
Accession Number: NP_001036.1
Length: 630 aa
Solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4)
Accession Number: NP_001036.1
Length: 630 aa
Analysis
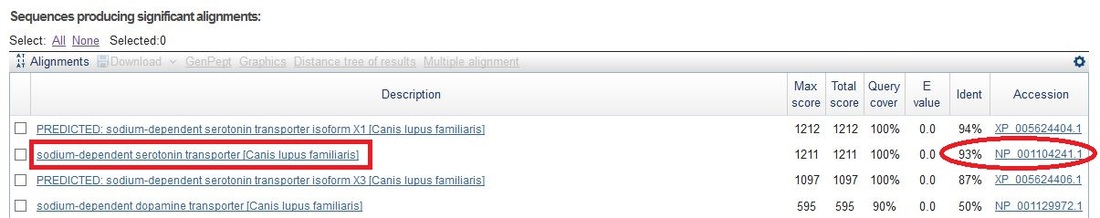
Using the accession number of SLC6A4, the NCBI Basic Local Alignment Search Tool (BLAST) was utilized to find the protein homologs of those organisms found to also be gene homologs. Since this study is on protein homologs, a protein blast was selected. A protein-protein BLAST algorithm was used to obtain the results. All seventeen organisms found to be gene homologs were also protein homologs. Each of their protein accession numbers and percent similarities were noted. For alignments that had predicted and validated sequences that were similar in their similarities, the validated sequence was chosen over the predicted for more accuracy.
Each of the seventeen organisms were confirmed by running a protein reciprocal BLAST, and some were also confirmed on HomoloGene. Each of the accession numbers from the original protein BLASTs were entered and searched within the homo sapiens database. If the human protein listed above came back as one of the top alignments, the organisms were considered protein homologs.
Results of the protein BLAST searches are as follows:
|
Pan troglodytes (Chimpanzee)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: XP_001135066.2 Max Identical: 99% Mus musculus (Mouse)
Solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 (Slc6a4) Accession Number: NP_034614.2 Max Identical: 93% Loxodonta africana (Elephant)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: XP_003416833.1 Max Identical: 93% Rattus norvegicus (Brown Rat)
solute carrier family 6 (neurotransmitter transporter), member 4 (Slc6a4) Accession Number: NP_037166.2 Max Identical: 92% Taeniopygia guttata (Zebra Finch)
solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 (SLC6A4) Accession Number: XP_004186287.1 Max Identical: 77% Drosophila melanogaster (Fruit Fly)
serotonin transporter (SerT) Accession Number: NP_523846.2 Max Identical: 55% |
Macaca mulatta (Rhesus Macaque)
solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 (SLC6A4) Accession Number: NP_001027995.1 Max Identical: 99% Equus ferus caballus (Domestic Horse)
serotonin transporter (5HTT) Accession Number: NP_001075293.1 Max Identical: 93% Bos Taurus (Cattle)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: NP_777034.1 Max Identical: 93% Columba livia (Dove)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: EMC77149.1 Max Identical: 78% Xenopus tropicalis (Frog)
sodium-dependent serotonin transporter-like Accession Number: XP_002932130.2 Max Identical: 71% Culex quinquefasciatus (Mosquito)
sodium-dependent serotonin transporter Accession Number: XP_001849951.1 Max Identical: 48% |
Felis catus (Cat)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: XP_006940067.1 Max Identical: 94% Canis lupus familiaris (Dog)
solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 (SLC6A4) Accession Number: NP_001104241.1 Max Identical: 93% Cricetulus griseus (Chinese Hamster)
solute carrier family 6 (neurotransmitter transporter), member 4 (Slc6a4) Accession Number: ERE67926.1 Max Identical: 92% Gallus gallus (Chicken)
solute carrier family 6 (neurotransmitter transporter), member 4 (SLC6A4) Accession Number: NP_998737.1 Max Identical: 77% Danio rerio (Zebrafish)
sodium-dependent serotonin transporter (slc6a4a) Accession Number: NP_001035061.1 Max Identical: 70% |
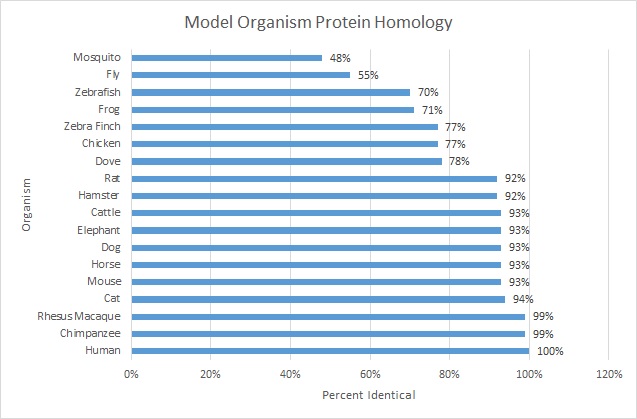
Figure 1: Graphical representation of protein homology, with most similar at the bottom and least similar at the top.
Discussion
As would be predicted based on the degree of conservation in genetic sequences between model organisms, there is a high amount of conservation between protein sequences as well. Non-animal bacteria and fungi are not represented in the data because SLC6A4 is not found outside of the kingdom Animalia. The protein homology does reveal that there is a higher degree of conservation between gene sequences than between protein sequences. One reason for this may be post-transcriptional modification, in which parts of the gene are changed or removed before translation occurs. The fact still remains that a relatively high proportion of the protein is maintained throughout evolution, from insects to humans. This indicates that SLC6A4 has some vital function for animals that is not necessary in non-animals.
References
[1] Homologies. (n.d.). Retrieved March 21, 2015, from http://evolution.berkeley.edu/evolibrary/article/lines_04
BLAST http://blast.ncbi.nlm.nih.gov/Blast.cgi?CMD=Web&PAGE_
HomoloGene http://www.ncbi.nlm.nih.gov/homologene
BLAST http://blast.ncbi.nlm.nih.gov/Blast.cgi?CMD=Web&PAGE_
HomoloGene http://www.ncbi.nlm.nih.gov/homologene